Base class for optimizer/sampler likelihood functions that compute likelihood at a point. More...
#include <Likelihood.h>
Public Member Functions | |
| int | getDataDim () const |
| Return the number of data points. More... | |
| int | getAmplitudeDim () const |
| Return the number of linear parameters (columns of the model matrix) More... | |
| int | getNonlinearDim () const |
| Return the number of nonlinear parameters (which parameterize the model matrix) More... | |
| int | getFixedDim () const |
| Return the number of fixed nonlinear parameters (set on Likelihood construction) More... | |
| ndarray::Array< Scalar const, 1, 1 > | getFixed () const |
| Return the vector of fixed nonlinear parameters. More... | |
| ndarray::Array< Pixel const, 1, 1 > | getData () const |
| Return the vector of weighted, scaled data points \(z\). More... | |
| ndarray::Array< Pixel const, 1, 1 > | getUnweightedData () const |
| Return the vector of unweighted data points \(y\). More... | |
| ndarray::Array< Pixel const, 1, 1 > | getWeights () const |
| Return the vector of weights \(w\) applied to data points and model matrix rows. More... | |
| ndarray::Array< Pixel const, 1, 1 > | getVariance () const |
| Return the vector of per-data-point variances. More... | |
| std::shared_ptr< Model > | getModel () const |
| Return an object that defines the model and its parameters. More... | |
| virtual void | computeModelMatrix (ndarray::Array< Pixel, 2,-1 > const &modelMatrix, ndarray::Array< Scalar const, 1, 1 > const &nonlinear, bool doApplyWeights=true) const =0 |
| Evaluate the model for the given vector of nonlinear parameters. More... | |
| virtual | ~Likelihood () |
| Likelihood (const Likelihood &)=delete | |
| Likelihood & | operator= (const Likelihood &)=delete |
| Likelihood (Likelihood &&)=delete | |
| Likelihood & | operator= (Likelihood &&)=delete |
Protected Member Functions | |
| Likelihood (std::shared_ptr< Model > model, ndarray::Array< Scalar const, 1, 1 > const &fixed) | |
Protected Attributes | |
| std::shared_ptr< Model > | _model |
| ndarray::Array< Scalar const, 1, 1 > | _fixed |
| ndarray::Array< Pixel, 1, 1 > | _data |
| ndarray::Array< Pixel, 1, 1 > | _unweightedData |
| ndarray::Array< Pixel, 1, 1 > | _variance |
| ndarray::Array< Pixel, 1, 1 > | _weights |
Detailed Description
Base class for optimizer/sampler likelihood functions that compute likelihood at a point.
Likelihood abstracts the problem of computing the likelihood over different kinds of data. It is responsible for creating a "model matrix" that maps amplitudes to data values, and maintaining a vector of scaled, weighted data values that corresponds to it. Its components can be represented best in the mathematical formula for a -log likelihood assuming Gaussian data and a model with both nonlinear parameters \(\theta\) and linear ("amplitude") parameters \(\alpha\):
\[ L(\alpha,\theta) = \frac{1}{2}\left(y - A(\theta)\alpha\right)^T\, \Sigma^{-1}\,\left(y - A(\theta)\alpha\right) \]
where \(y\) is the data vector, \(\Sigma\) is the data covariance matrix (assumed to be diagonal), and \(A(\theta)\) is the "true" model matrix (parametrized on the nonlinear parameters).
When fitting or sampling from the likelihood, however, we don't want to use these quantities directly, and they aren't what the Likelihood class provides. Instead, we reparametrize with:
\[ w_i \equiv \Sigma_{i,i}^{-1/2} \]
\[ z_i = w_i y_i \]
\[ B_{i,j} = w_i A_{i,j} \]
resulting in the equivalent formula:
\[ L(\alpha,\theta) = \frac{1}{2}\left(z-B(\theta)\alpha\right)^T\,\left(z-B(\theta)\alpha\right) \]
The \(w_i\) are the weights, which are applied to both the data vector and the model matrix to account for the noise in the data. In some cases, we may choose to use a constant weight rather than per-pixel weights, but will will still use a vector to represent it.
Definition at line 69 of file Likelihood.h.
Constructor & Destructor Documentation
◆ ~Likelihood()
|
inlinevirtual |
Definition at line 126 of file Likelihood.h.
◆ Likelihood() [1/3]
|
delete |
◆ Likelihood() [2/3]
|
delete |
◆ Likelihood() [3/3]
|
inlineprotected |
Definition at line 138 of file Likelihood.h.
Member Function Documentation
◆ computeModelMatrix()
|
pure virtual |
Evaluate the model for the given vector of nonlinear parameters.
- Parameters
-
[out] modelMatrix The dataDim x amplitudeDim matrix \(B\) that expresses the model projected in such a way that it can be compared to the data when multiplied by an amplitude vector \(\alpha\). It should be weighted if the data vector is. The caller is responsible for guaranteeing that the shape of the matrix correct, but implementations should not assume anything about the initial values of the matrix elements. [in] nonlinear Vector of nonlinear parameters at which to evaluate the model. [in] doApplyWeights If False, do not apply the weights to the modelMatrix.
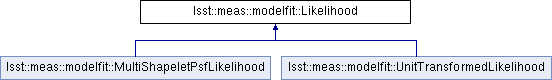
Implemented in lsst::meas::modelfit::UnitTransformedLikelihood, and lsst::meas::modelfit::MultiShapeletPsfLikelihood.
◆ getAmplitudeDim()
|
inline |
Return the number of linear parameters (columns of the model matrix)
Definition at line 77 of file Likelihood.h.
◆ getData()
|
inline |
Return the vector of weighted, scaled data points \(z\).
Definition at line 89 of file Likelihood.h.
◆ getDataDim()
|
inline |
◆ getFixed()
|
inline |
◆ getFixedDim()
|
inline |
Return the number of fixed nonlinear parameters (set on Likelihood construction)
Definition at line 83 of file Likelihood.h.
◆ getModel()
|
inline |
Return an object that defines the model and its parameters.
Definition at line 105 of file Likelihood.h.
◆ getNonlinearDim()
|
inline |
Return the number of nonlinear parameters (which parameterize the model matrix)
Definition at line 80 of file Likelihood.h.
◆ getUnweightedData()
|
inline |
Return the vector of unweighted data points \(y\).
Definition at line 92 of file Likelihood.h.
◆ getVariance()
|
inline |
Return the vector of per-data-point variances.
Definition at line 102 of file Likelihood.h.
◆ getWeights()
|
inline |
Return the vector of weights \(w\) applied to data points and model matrix rows.
Will be an empty array if no weights are applied.
Definition at line 99 of file Likelihood.h.
◆ operator=() [1/2]
|
delete |
◆ operator=() [2/2]
|
delete |
Member Data Documentation
◆ _data
|
protected |
Definition at line 149 of file Likelihood.h.
◆ _fixed
|
protected |
Definition at line 148 of file Likelihood.h.
◆ _model
|
protected |
Definition at line 147 of file Likelihood.h.
◆ _unweightedData
|
protected |
Definition at line 150 of file Likelihood.h.
◆ _variance
|
protected |
Definition at line 151 of file Likelihood.h.
◆ _weights
|
protected |
Definition at line 152 of file Likelihood.h.
The documentation for this class was generated from the following file:
- /j/snowflake/release/lsstsw/stack/lsst-scipipe-0.7.0/Linux64/meas_modelfit/22.0.1-4-g44f2e3d+9e4ab0f4fa/include/lsst/meas/modelfit/Likelihood.h